Genomic Unity® 2.0 Case Study
Overview
Patient:
2-year-old male
Clinical presentation:
Global developmental delays, exophoria, highly elevated alkaline phosphatase, severe congenital hypotonia, strabismus, multiple phenotypic anomalies including relative macrocephaly, high forehead, pectus excavatum and more
Testing strategy:
Variantyx whole genome testing
Key finding:
De novo, heterozygous, pathogenic 10.54 kb deletion in the GRIA2 gene
Clinical outcome:
Diagnosis established
Why Genomic Unity® 2.0 was the right choice
Despite a prior negative exome, the patient’s severe hypotonia, exophoria, developmental delays and distinctive features were strongly suggestive of an underlying genetic condition. When a highly elevated alkaline phosphatase level was identified, suspicion grew for a hyperphosphatasia with impaired intellectual development syndrome, despite the absence of classical physical findings.
Genomic Unity® 2.0 was selected because it delivers the most comprehensive genomic insight from the start while:
- Reducing time to diagnosis
- Avoiding unnecessary testing
- Supporting the highest standard of patient care
Diagnostic finding: GRIA2-related neurodevelopmental disorder
Variantyx Genomic Unity® 2.0 testing identified a de novo, heterozygous, pathogenic 10.54 kb deletion in the GRIA2 gene.
Encompassing the 5′ UTR and a portion of exon 1, the deletion is expected to result in loss of protein function.
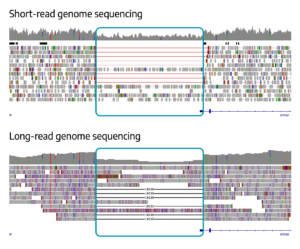
Uniform data from both short-read WGS (top) and long-read WGS (bottom) clearly show the deletion, with long-read WGS providing precise sizing of 10,539 bp.
Impact on clinical care
Established an elusive diagnosis.
Variant spotlight: Partial exon deletion
Detection challenges:
Partial exon deletions are below the limit of detection of most genetic tests – including exome and panel tests. Especially when one or both breakpoints fall within an intronic or intergenic region. This leads to missed variant calls.
Why Genomic Unity® 2.0
- Has a detection range from 1 bp to whole chromosomal events, easily detecting partial exon deletions.
- Sequences all intronic and intergenic regions, enabling breakpoint detection regardless of location.
Additional similar cases
Genomic Unity® – Partial exon deletion explains clinical Rett syndrome diagnosis
Genomic Unity® – Partial exon deletion plus deep intronic SNV explains juvenile parkinsonism
IriSight® – Partial DOK7 deletion plus small indel explains multiple fetal anomalies
Variantyx tests that would have identified this variant
Genomic Unity® 2.0 | Genomic Unity® Whole Genome Analysis | Genomic Unity® Lightning Genome Analysis | Genomic Unity® Lightning 2.0 Genome Analysis – NICU | Genomic Unity® Lightning 2.0 Genome Analysis – Standard | Genomic Unity® Exome Plus Analysis | Genomic Unity® Exome Analysis | Genomic Unity® Constitutional Genome-Wide Copy Number Variant Analysis | Genomic Unity® Genome-Wide CNV and FMR1 Analysis